Content uploaded by Trudi Schüpbach
Author content
All content in this area was uploaded by Trudi Schüpbach on Apr 02, 2018
Content may be subject to copyright.
ELSEVIER Mechanisms of Development 59 (1996) 105-113
The Drosophila TGF-a-like protein Gurken: expression and
cellular localization during Drosophila oogenesis
F. Shira Neuman-Silberbergl, Trudi Schi.ipbach*
HHMI, Department of Molecular Biology, Princeton University, Princeton, NJ 08544, USA
Received 27 March 19%; revision received 11 June 1996; accepted 11 June 1996
Abstract
The establishment of anterior-posterior and dorsal-ventral polarity of the Drosophila egg and embryo depends on the function of the
genes gurken, comichon and Egf? (Drosophila epidermal growth factor receptor homolog). These genes encode components of a signal
transduction pathway that transmits information between the germline cells and the somatic follicle cells of the ovary. gurken encodes a
transforming growth factor-a-like protein and is a putative germline ligand of the Egfr present on the follicle cells. In mid-oogenesis
the gurken transcript becomes spatially localized to the future dorsal-anterior cortex of the oocyte. To analyze the distribution pattern of
Gurken protein we prepared antibodies against Gurken. We describe here the distribution pattern of the Gurken protein in wild-type
ovaries and in ovaries from a number of dorsal-ventral patterning mutants. By immunoblotting we detect one major form of the Gurken
protein, which likely corresponds to the unprocessed protein.
Keywords: Drosophila; Dorsal-ventral axis; Oogenesis; Signal transduction; Confocal
1. Introduction
In Drosophila the information required for the estab-
lishment of both body axes (anterior-posterior and dorsal-
ventral) in the egg and subsequently in the embryo is in-
corporated into the egg chamber during oogenesis (for
review of oogenesis, see Spradling, 1993). The establish-
ment of axial polarity requires the function of a number
of genes, among them the genes gurken (grk), Egfr (also
designated torpedo (top) or DER) and cornichon (cni)
(Schtipbach, 1987; Schtipbach et al., 1991; Gonzales-
Reyes et al., 1995; Roth et al., 1995). Females mutant for
any of these genes produce ventralized eggs and embryos,
where the ventral cell fates are expanded and dorsal cell
fates are reduced (Schtipbach, 1987; Schiipbach et al.,
1991; Roth et al., 1995). In addition, females harboring
the most severe allele combinations at these loci produce
egg chambers with a duplication of the anterior follicle
* Corresponding author.
’ Present address: Department of Molecular Microbiology and Bio-
technology, Tel-Aviv University, 69978 Tel Aviv, Israel.
cell fates at the posterior end (Gonzalez-Reyes et al.,
1995; Roth et al., 1995).
grk, Egfr (top/DER) and cni are components of a sig-
nal transduction pathway that provides communication
between the germline and the somatic follicle cells of the
ovary. The function of Egfr, which encodes the Droso-
phila epidermal growth factor receptor homolog (Price et
al., 1989; Schejter and Shilo, 1989), is required in the
follicle cells, whereas the functions of grk and cni are
required in the oocyte (Schupbach, 1987; Schiipbach et
al., 1991). grk encodes a transforming growth factor
(TGF)-a-like protein that may serve as a germline ligand
of the Egfr (Neuman-Silberberg and Schiipbach, 1993).
cni encodes a novel protein (Roth et al., 1995). The es-
tablishment of posterior follicle cell fate, early in
oogenesis, depends on the transduction of a signal from
the oocyte to the follicle cells via Egfr. In turn, the newly
determined posterior follicle cells send a signal to the
oocyte that leads to rearrangement of the cytoskeleton
(Ruohola et al., 1991; Gonzalez-Reyes et al., 1995; Roth
et al., 1995). This cytoskeletal rearrangement is required
for the proper localization of determinants to the anterior
0925-4773/96/$15.00 0 1996 Elsevier Science Ireland Ltd. All rights reserved
,.,r nrrnrr l-“-l,.I\,.,.rl” _
106 F. Shira Neumun-Silberberg, T. Schiipbach /Mechanisms of Development 59 (1996) 10.5-113
and posterior poles (Pokrywka and Stephenson, 1991;
Clark et al., 1994; Lane and Kalderon, 1994) and hence
for the establishment of anterior-posterior, polarity (for
reviews see St. Johnson, 1995; Rongo and Lehmann,
1996).
Dorsal-ventral polarity in the egg chamber is first ap-
parent during mid stages of oogenesis, when the oocyte
nucleus, which is positioned in the center of the oocyte,
relocates to the future dorsal-anterior corner of the oocyte
(for review see Mahowald and Kambysellis, 1980). This
movement of the oocyte nucleus depends also on the prior
rearrangement of the microtubule network (Koch and
Spitzer, 1983; Gonzalez-Reyes et al., 1995; Roth et al.,
1995). Immediately after the nucleus moves, the grk RNA
present in the oocyte becomes localized to the future dor-
sal-anterior corner of the oocyte in close proximity to the
oocyte nucleus (Neuman-Silberberg and Schtipbach,
1993).
Spatial restriction of the grk transcript to the future
dorsal-anterior corner of the oocyte is essential for correct
dorsal-ventral patterning. Increasing the grk gene dosage
results in partially mislocalized grk RNA and dorsaliza-
tion of eggs and embryos (Neuman-Silberberg and
Schupbach, 1994). Females mutant for the genes squid
(sqd), fs(l)KlO, cappuccino (cupu) and spire (spir) also
produce dorsalized egg chambers (Wieschaus et al., 1978;
Wieschaus, 1979; Manseau and Schiipbach, 1989; Schilp-
bath et al., 1991; Kelley, 1993). In such egg chambers the
grk transcript is mis-localized and is found as an anterior
ring around the entire circumference of the egg chamber
(Neuman-Silberberg and Schiipbach, 1993). These obser-
vations suggest that grk encodes a spatially restricted li-
gand that acts to locally activate Egfr, thus defining the
dorsal side of the egg chamber. To determine whether the
pattern of the grk mRNA reflects the distribution of the
Grk protein, we have prepared antibodies against the Grk
protein. Using these antibodies we have identified the Grk
protein in ovarian extracts and characterized its expres-
sion pattern in egg chambers produced by wild-type fe-
males and by females mutant for genes in the grk/Egfr
signal transduction pathway. Our results imply a dynamic
pattern for the Grk protein throughout oogenesis and fur-
ther illustrate the need for spatial restriction of the Grk
protein in dorsal-ventral patterning.
2. Results
2.1. Antibodies against a GST-GRK fusion protein
recognize a 46 kDa protein present in ovarian extracts
Polyclonal antibodies were prepared against a 44 kDa
fusion protein containing 133 amino acid residues (53-
185) from the extra-cellular domain of the Gurken (Grk)
protein (Fig. 1A) fused to glutathione-S-transferase
(GST). The fusion protein was partially purified by ab-
sorption to glutathione-coupled beads and injected into
rats. The antiserum was tested for its ability to detect Grk
protein by immunoblotting of extracts prepared from
ovaries and adult males (Fig 1B). This antiserum recog-
nizes a 46 kDa protein present in ovaries from wild-type
females (lane 1) but not in males (data not shown). The
46 kDa band very likely represents the Grk protein since a
protein of an apparently similar molecular weight is pro-
duced in an in vitro translation system (Fig. lC, lane 2).
Furthermore, the level of this protein is dramatically di-
minished in ovaries from a number of grk mutants (grkHK,
grkHF, grkzB6) (Fig. lB, lanes 24). In other mutants in the
grWEgfr signaling pathway (Egfr, cni, sqd, cupu), the size
of the protein and the total level of the protein is roughly
similar to that observed in wild-type (data not shown). An
additional gene, orb, was also shown to be required for
grk RNA localization during oogenesis (Christerson and
McKearin, 1994; Roth and Schtipbach, 1994). In ovaries
from a hypomorphic orb combination (orbme1/orbF343) the
A
SP EGF
129
14kdWltW~
4 tub
Fig. I. Antibodies prepared against the Grk protein recognize a 46 kDa
protein in ovarian extracts. (A) A 400 bp Sphl-Hind3 fragment from the
grk cDNA corresponding to amino acid residues 53-185 (133 amino
acids) of the extra cellular domain of Grk was subcloned into the PGEX
expression vector (Promega) and the resulting fusion protein was used
to immunize rats. (B) Extracts prepared from ovaries and from whole
male flies were fractionated by polyacrylamide gel electrophoresis and
blotted onto a nitrocellulose filter. Blots were probed with either the
anti-Grk antibody (dilution 1:4000) or with an anti-a-tubulin antibod-
ies Lane 1, ovarian extract from wild-type females. Lanes 2, 3 and 4,
ovarian extracts from grk females (grkHK, grkHF, and grkzB, respec-
tively). Lane 5, ovarian extracts from orb females. (C) Protein synthe-
sized in vitro using the grk cDNA as a template. Lane 1, ovarian ex-
tracts from wt females. Lanes 2 and 3, in vitro translation products in
the TNT T7 transcription/translation system with (2) or without (3) the
I .7 grk cDNA plasmid as template. The antibodies recognize an addi-
tional less abundant protein of 76 kDa, whose level is not affected by
the grk mutations (see Section 4). In the experiment involving the in
vitro translation system the Grk protein appeared as a doublet (C).
However, in all other preparations the protein always appeared as a
single band.
F. Shira Neumun-Silberberg, T. Schiipbach / Mechanisms of Development 59 (1996) 105-I 13 107
Grk protein level appears to be much lower than in wild-
type (Fig lB, lane 5).
2.2. The Grk protein is spatially restricted to the dorsal
anterior corner of the oocyte, closely resembling the RNA
localization pattern
Using the anti Grk antiserum for whole mount ovary
staining, we found that the Grk protein is already ex-
pressed in early stages of oogenesis. We first detect the
protein in region 2B of the germarium (data not shown)
which coincides with the earliest detection of the grk tran-
script (Neuman-Silberberg and Schtipbach, 1993). Similar
to the RNA, the protein is localized to the oocyte in
young egg chambers (Fig. 2A), where it appears as
granular staining (Fig 2B). We have previously shown
that during stage 8 of oogenesis, after the oocyte nucleus
assumes an asymmetric position at the future dorsal-
anterior corner of the oocyte, the grk transcript becomes
localized to the dorsal-anterior cortex of the oocyte, in
close proximity to the oocyte nucleus (Neuman-
Silberberg and Schiipbach, 1993). Likewise, the Grk
protein becomes spatially restricted in mid-stages of
oogenesis and its distribution closely resembles the pat-
tern observed for the RNA. The protein is exclusively
found at the future dorsal cortex of the oocyte where it
co-localizes with the membrane associated F-actin (Fig.
2C,D). We did not observe any punctate or granular ac-
cumulation of Grk protein in the intercellular space be-
tween oocyte and follicle cells, that would resemble e.g.
the pattern for secreted Wingless protein (Bejsovec and
Wieschaus, 1995). At later stages of oogenesis (10-12)
the distribution of the Grk protein is different from that of
the grk RNA. While the grk RNA is still tightly localized
to the dorsal-anterior cortex of the oocyte, the Grk protein
forms an elongated anterior to posterior stripe extending
over approximately half the length of the oocyte’s dorsal
midline (Fig. 2E,F). This is the one stage in egg chamber
development where the protein distribution cannot be
precisely predicted by the RNA localization pattern.
2.3. Grk protein is mislocalized in egg chambers from
females mutant for the genes fs(l)KIO, sqd, capu ana’ orb
which are required for proper localization of the grk
RNA.
We have examined the expression pattern of the Grk
protein in egg chambers from fs(l)KlO, sqd and capu
females that produce dorsalized egg chambers. The grk
transcript in these egg chambers is mis-localized and
Fig. 2. Distribution pattern of the Grk protein in wild-type egg cham-
bers. Confocal images of indirect immunofluorescent staining of egg
chambers with anti-Grk antibody using a CY3conjugated anti-rat sec-
ondary antibody. The Grk protein is shown in green and F-actin is
visualized with rhodamine-conjugated phalloidin in red; regions where
labels overlap are shown in yellow. (A) Young egg chambers stage 2-
5; Grk is localized to the oocyte. (B) Stage 6 egg chamber; Grk protein
appears as granules in the ooplasm and at the cortex of the oocyte.
(C) Stage 8 egg chamber; Grk protein is localized to the anterior-dorsal
cortex of the oocyte. (D) Stage 9 egg chamber. (E) Late stage IOA egg
chamber. (F) Stage IOB egg chamber. Grk appears as an elongated
stripe along the dorsal cortex. Scale bar: (A,D,E) approximately 32pm;
(B) approximately 25pm; (CF) approximately 38,~m. In (A,B) the
oocyte nucleus is at the posterior pole; in (B-F) the oocyte nucleus is at
the dorsal anterior comer of the oocyte.
108 F. Shira Neuman-Silberberg, T. Schiipbach /Mechanisms of Development 59 (I 996) 105-l 13
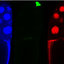
Fig. 3. Distribution pattern of Grk protein in egg chambers produced by females mutant forfs(Z)HO, sqd, capu and orb. (A) Egg chamber from a wild-
type female. (B) Egg chamber from fs(l)KIO mutant. (C) Egg chamber from capu mutant. (D) Egg chamber from sqd mutant. All egg chambers are at
stage 9 of oogenesis. In egg chambers produced by the mutants, the Grk protein is not limited to the dorsal-anterior cortex of the oocyte as in wild-
type. A certain fraction of the protein seems to be correctly localized; however, significant levels of the protein reside at the anterior margin of the egg
chamber around the entire circumference. (E,F) Egg chambers mutant for orb, stages 9 and lOB, respectively. In orb egg chambers the Grk protein is
more symmetrically localized to the oocyte cortex. Labels and colors are as indicated in Fig. 2. Scale bar, approximately 25 pm. The position of the
oocyte nucleus is indicated by a white asterisk.
Fig. 4. Grk protein pattern in egg chambers produced by a number of grk, Egfr and cni mutants. (A) Egg chamber mutant for grpc/grkDc. Although
most of the Grk protein is correctly localized a considerable amount of the protein appears as dispersed granules in the ooplasm of sta e 8 and older
egg chambers. (B) Egg chamber from grk 7a5/grkHK. Grk protein appears as an aggregate in the ooplasm. (C) Egg chamber from grgE* t gdE*‘. The
oocyte nucleus stayed at the posterior end. Some of the Grk protein is localized to the posterior cortex while the rest a pears as granular staining in the
ooplasm. (D) Egg chamber from cn~“2/cniiAR5S. Staining similar to grsE”. (E) Egg chamber from Egfrco&fii & . In this late stage 9 egg cham-
ber the oocyte nucleus failed to assume an anterior dorsal position and stayed at the posterior end. The Grk protein also resides at the posterior cortex
in close proximity to the oocyte nucleus. All egg chambers are at stage 9 of oogenesis: (A,B,D) early stage 9; (C,E) late stage 9. Labels and colors are
as indicated in Fig. 2. Scale bar: (A,C-E) approximately 25 pm; (B) approximately 28 pm. The position of the oocyte nucleus is indicated by a white
asterisk.
F. Shiru Neuman-Silberberg, T. Schiipbach / Mechanisms uf Development 59 (1996) 105-I 13 109
resides at the anterior margin of the oocyte around the
entire circumference of the egg chamber (Neuman-
Silberberg and Schtipbach, 1993). In vitellogenic egg
chambers (past stage 7) produced by these mutants, the
Grk protein is also found at the anterior margin of the
oocyte all around the circumference, in a pattern similar
to that of the grk transcript (Fig. 3B-D) (forfi(l)KZO see
also Serano et al., 1995). In addition, a significant amount
of the protein is associated with the asymmetrically posi-
tioned oocyte nucleus, like in wild-type. This pattern re-
sembles again the pattern observed for the grk RNA.
Females mutant for the gene orb produce a variety of
defective egg chambers, among them a certain fraction of
‘lateralized’ egg chambers with no clear dorsal-ventral
polarity (Christerson and McKearin, 1994; Roth and
Schupbach, 1994). In such egg chambers the grk tran-
script appears to be more evenly distributed in the oocyte
cytoplasm (Roth and Schiipbach, 1994). In young egg
chambers produced by orb females the Grk protein is
normally localized to the oocyte as in wild-type. Many of
the older egg chambers seem to have low levels of the
Grk protein. In the few egg chambers in which the Grk
protein is detectable by whole mount antibody staining,
the Grk protein is evenly distributed around the oocyte
cortex rather than being preferentially associated with the
future dorsal cortex of the oocyte (Fig. 3E,F). Similar
results were also obtained by Jacqueline Chang and Paul
Schedl (unpublished).
2.4. Expression and distribution of the Grkprotein in egg
chambers produced by grk, Egfr and cni mutants
By whole mount ovary staining we could distinguish
three classes of grk mutants based on the expression and
distribution pattern of the Grk protein. In the first class,
protein distribution is very similar to the pattern observed
in wild-type (grkwG, grkm7, grkED22). In the second class,
low levels of RNA are detected (Neuman-Silberberg and
Schtipbach, 1993) and the level of protein present is be-
low the level of detection when staining whole ovaries
(grk2B6, grkEDtl, grkHF, grkHK, grkHL, grkHG). In the third
class, the level of Grk protein is comparable to the protein
level in wild-type ovaries. Nevertheless, in mid-stages of
oogenesis only a fraction of the protein is found localized
to the future dorsal-anterior cortex of the oocyte. The rest
of the protein appears as granular staining in the ooplasm
(grk2E*2, grkQt, grkDc, grk705) (see Fig. 4). Such cyto-
plasmic staining is found in wild-type only in pre-
vitellogenic egg chambers (prior to stage 7). In most egg
chambers produced by Egfr females the Grk protein is
normally localized to the dorsal cortex of the oocyte (see,
however, Fig. 4E for an egg chamber in which the oocyte
nucleus and the Grk protein are mispositioned at the pos-
terior pole). Similar to some of the grk mutants, there is
granular protein staining in the ooplasm of vitellogenic
egg chambers from cni mutants (Fig. 4D).
In a fraction of egg chambers produced by females
with a strong combination of cni alleles (cniAR55/cniAA’2)
or in severe allele combinations of grk and Egfr mutants,
the oocyte nucleus does not move to the dorsal-anterior
corner of the oocyte but stays at the posterior end
(Gonzales-Reyes et al., 1995; Roth et al., 1995). In this
case the grk RNA resides at the posterior in close prox-
imity to the oocyte nucleus and the corresponding protein
appears also to be localized to the posterior cortex (Fig.
4C-E).
3. Discussion
3.1. Identification of the Grk protein in ovarian extracts
From sequence analysis, the grk gene is predicted to
encode a 33 kDa protein (Neuman-Silberberg and Schtip-
bath, 1993). Using an antibody made against a GST fu-
sion protein containing most of the Grk extra-cellular
domain (excluding the EGF repeat), we detect a predomi-
nant protein of an apparent molecular weight of 46 kDa.
The amount of this protein is reduced in ovaries from
several of the grk mutants. Furthermore, a protein which
co-migrates with the 46 kDa band is synthesized in a
coupled in vitro transcription/translation system only
when grk cDNA is included in the reaction mixture.
These data indicate that the 46 kDa protein detected by
immunoblotting most likely represents the unmodified
form of the Grk protein. As mentioned in the legends to
Fig. 1, we also detect a weaker band of 76 kDa. This
76 kDa protein is generally present even in the mutants
that have reduced levels of the 46 kDa protein and, there-
fore, is probably not related to the Grk protein (see Sec-
tion 4). Smaller polypeptides, that may indicate process-
ing of Grk, similar to the vertebrate TGF-a family of
growth factors, were not detected in extracts from wild-
type ovaries using an antibody directed against the EGF-
repeat of Grk (unpublished).
3.2. The Grk protein exhibits a dynamic pattern that
changes throughout egg chamber development
The Grk protein is expressed in the oocyte early in
oogenesis and is apparent as soon as the RNA is detect-
able, in region 2B of the germarium. In pre-vitellogenic
egg chambers a small fraction of the Grk protein colocal-
izes with the membrane-associated F-actin, whereas most
of the Grk protein appears as granular staining in the
ooplasm. Since from its sequence Grk is predicted to be
an exported protein, this granular staining may reflect
association with the rough endoplasmic reticulum and
possibly with other components of the cellular secretory
machinery. After the grk RNA becomes localized to the
dorsal-anterior cortex, Grk protein disappears from the
ooplasm and is exclusively found at the future dorsal
cortex of the oocyte. Grk is likely to function as an extra-
110 F. Shiru Neumun-Silberberg, T. Schiipbach /Mechanisms #Development 59 (1996) 105-I 13
cellular ligand, and its sequence predicts the presence of a
transmembrane domain. It is, therefore, reasonable to
assume that the cortical localization of Grk reflects its
association with the oocyte’s membrane. Regulating the
levels of Grk protein in the membrane of young egg
chambers and spatially restricting Grk to the dorsal-
anterior corner of the oocyte in mid-oogenesis could be
achieved by several different mechanisms. One possibility
is that in young oocytes the overall organization of the
cytoskeleton may be different from the organization in
later stage oocytes. Since transport vesicles move along
the cytoskeleton, a less organized cytoskeleton at earlier
stages might lead to the accumulation of vesicles in the
cytoplasm of these stages. A dependence of Grk protein
localization on the cytoskeletal architecture may be sug-
gested by our finding that in several of the grk and cni
mutants various amounts of granular staining of the Grk
protein is apparent in the ooplasm of vitellogenic egg
chambers. In some cases the protein appears as a dense
aggregate in the ooplasm. These clumps are reminiscent
of the accumulation pattern observed for the plus-end-
directed microtubule motor protein kinesin in egg cham-
bers from severe cni, grk and Egfr mutant females. In
such mutant egg chambers a kinesin-/I-galactosidase fu-
sion protein accumulates at the center of the ooplasm
rather than at the posterior end (Gonzales-Reyes et al.,
1995; Roth et al., 1995).
Another interesting point is the distribution pattern of
the Grk protein in older egg chambers (stages 10-l 1). At
these stages the Grk protein forms an elongated dorsal
cortical stripe extending along almost half the oocyte’s
length. This is in contrast to the grk RNA that remains
relatively tightly localized to the dorsal-anterior corner of
the oocyte, in close proximity to the oocyte nucleus. Dif-
ferent mechanisms could account for this protein pattern.
One possibility is that the protein diffuses in the oocyte
membrane in a spatially restricted manner forming a
stripe. Alternatively, the oocyte membrane may be
growing at the anterior end, displacing the older Grk
protein posteriorly. In any case, the Gurken protein re-
mains asymmetrically distributed within the oocyte mem-
brane over a considerable length of time (the time period
from stage 8 to stage 10B comprises about 34 h; Sprad-
ling, 1993). It is likely that some localization or anchoring
mechanism operates on the transmembrane form of the
protein to prevent it from diffusing and spreading within
the oocyte membrane.
The Grk protein appears localized to the dorsal side of
the oocyte at stage 8. Since the follicle cells will migrate,
or condense, over the oocyte during stage 9, a larger
group of lateral follicle cells will be exposed to high lev-
els of Grk protein as they migrate past the region of the
oocyte where Grk is concentrated. It is therefore possible
that the follicle cell migration results in an induction of
dorsal follicle cell fates in a larger group of follicle cells
than the group that is seen overlying the Grk protein in
any one static picture. However, expression of rhomboid
RNA in dorsal follicle cells, which may be one of the first
responses to grtVEgfr signaling, does not extend very far
posteriorly when it is first detectable in late stage 9 of
oogenesis. Consequently, at least rhomboid expression
does not seem to be stably induced in the follicle cells that
were transiently exposed to high levels of Grk protein
when migrating over the oocyte during the earliest stages
of follicle cell migration (Ruohola-Baker et al., 1993;
Neuman-Silberberg and Schiipbach, 1994).
Grk protein persists long past stage 9 of oogenesis,
which is presumably the point when dorsal-ventral polar-
ity is determined in the egg chamber (Monte11 et al.,
1991). It is appealing to speculate that Grk protein may be
required both for the establishment of dorsal follicle cell
fate as well as for the maintenance of this differentiated
state. On the other hand, it is possible that Grk protein has
a very long half life and is, therefore, persistent in the
oocyte many hours after it has fulfilled its function. The
generation of conditional grk alleles may enable us to
address this issue.
3.3. Spatial restriction of the Grk protein is required for
proper dorsal-ventral patterning
We have previously shown that grk RNA is mislocal-
ized in dorsalized egg chambers produced by fs(l)K10,
sqd and capu females. This result strongly suggested that
the grk RNA residing around the anterior margin may be
responsible for the dorsalization of the egg chambers in
such mutants. This point is further supported by the ob-
servation that the Grk protein follows the RNA pattern
and is also present at the anterior margin around the entire
circumference in dorsalized egg chambers. Therefore, the
spatial restriction of the Grk protein to the future dorsal
side of the oocyte provides the initial molecular informa-
tion for the establishment of an asymmetric follicular
epithelium.
In the case of orb mutants, we have shown here that
the level of Grk protein is reduced in these egg chambers
and that the protein is more evenly distributed in the oo-
cyte cortex rather than being restricted to the future dorsal
side. Such a distribution pattern of the Grk protein may
account for the presence of partially lateralized eggs with
no obvious dorsal structures among eggs laid by orb fe-
males. Unlike the dorsalization observed for fs(l)KlO,
sqd and capu mutants, which have normal levels of the
Grk protein, the levels of Grk in egg chambers produced
by orb mutants may not be sufficient to cause a dorsalized
phenotype. It has been recently shown that the Xenopus
homolog of orb, CPEB, functions in polyadenylation of
mRNAs in the Xenopus oocyte (Hake and Richter, 1994).
In addition, polyadenylation of specific Drosophila ma-
ternal mRNAs was found to regulate their translational
activation (Salles et al., 1994). The orb product may also
function in translational regulation of mRNAs during
F. Shim Neumun-Silberberg, T. Schiipbuch / Mechunisms of Development 59 (1996) 105-I 13 111
Drosophila oogenesis, which would account for the low
levels of the Grk protein found in orb mutants. A model
describing a role of Orb in tethering of RNAs and on-site
translation has been proposed by Christerson et al. (1995).
4. Experimental procedures
4. I. Fly stocks
Oregon R (Lindsley and Grell, 1968) was used as
wild-type. Thirteen ethyl methanesulfonate- induced al-
leles of grk were used: grkwG, grkHK, grkHG, grkHL , grkQ’,
grkDC, grkHF (Schiipbach, 1987; Neuman-Silberberg and
Schupbach, 1993); grkZB6, grkZE12, grk70s (induced on a b
pr cn sea chromosome; T.S. and D. St. Johnston, unpub-
lished data); grkED22, grkm7, grkED” (Clifford and
Schupbach, 1989). Additional strains used were: fs(l)KlO
(Wieschaus et al., 1978); EgfrQn, Egfr” (Schiipbach,
1987); Egfrco (Clifford and Schiipbach, 1989); cniARs5 ,
cniAA12 (Roth et al , 1995); sqdAN240 (Kelley, 1993);
capuRK12 (Manseau and Schiipbach, 1989); orbme’, orbF343
(Lantz et al., 1992; Christerson and McKearin, 1994). To
obtain the most severe phenotypes for E&r, cni and sqd,
females carrying these mutations in trans to deficiencies
Df(2R)topJXA (Price et al., 1989), Dfl2L)Zlll8 (Roth et al.,
1995) and Df(3L)urd (Kelley, 1993), respectively, were
constructed. All mutant alleles were kept as balanced
stocks. For marker mutations and deficiencies see Lind-
sley and Zimm (1992).
4.2. Antibody preparation
A 400 bp SphI-Hind3 fragment from the grk cDNA
was subcloned into the PGEX expression vector
(Promega). Expression of this construct in bacteria gives
rise to a 44 kDa fusion protein that included 133 amino
acid residues (53-185) from the extracellular domain of
the Grk protein fused in frame downstream of the GST
protein. The fusion protein was partially purified by ab-
sorption to glutathione-coupled agarose beads (Sigma)
and used for the production of polyclonal antiserum in
rats (Pocono Rabbit Farm and Laboratory).
4.3. Specificity of the antiserum
Originally, several rats were immunized with the same
fusion protein. In the initial tests all sera detected the fu-
sion protein. On Western blots of ovarian extracts all sera
detected multiple bands. We determined that rat no. 10
yielded a serum that reacted with the smallest number of
protein bands, and we therefore used the antiserum from
this rat for our experiments. However, we also confirmed
the conclusions regarding the whole mount staining pat-
terns using the serum from the other rats.
The specificity of the preabsorbed antiserum was also
tested on ovarian extracts from females homozy’gous
mutant for grkHK and grk2B. These females produce very
little, if any, grk RNA (Neuman-Silberberg and Schtip-
bath, 1993). In whole mount preparations of egg cham-
bers from these females the preabsorbed serum did not
produce any specific staining, and we conclude, therefore,
that on whole mounts the serum detects predominantly
the Gurken protein. On Western blots from ovarian ex-
tracts from the same mutant females immunoreactivity
with a 46 kDa band is strongly reduced compared to ex-
tracts from wild-type females, and this band therefore
corresponds to the Gurken protein. The antiserum still
recognizes a band of about 76 kDa; however, this band
was not consistently observed in all samples, and was
sometimes barely detectable, even in extracts from wild-
type females. We therefore interpret the 76 kDa band as a
cross-reacting protein that is probably not related to Gur-
ken.
4.4. Whole-mount antibody staining
Ovaries were fixed for 20 min in 4% paraformalde-
hyde saturated with heptane including 0.5% NP40. The
antibody staining procedure was performed as described
by Peifer et al. (1993). Prior to use, the anti Grk antibody
was pre-absorbed using fixed ovaries from grkHK mutant
females that have reduced levels of the Grk product. The
primary antibody was used at a final dilution of 1:3000.
Cy3-conjugated goat anti-rat (Jackson Immunoresearch
Laboratories) (pre-absorbed with wild-type ovaries) at a
final dilution of 1: 1000 was used as a secondary antibody.
FITC-labeled Phalloidin (Sigma) was used as a counter
stain. Stained ovaries were mounted in Aqua-polymount
(Polysciences, Inc.) and visualized on a BioRad confocal
microscope.
4.5. Biochemical methods
Protein extracts were prepared from males and from
ovaries of 4-5 days old females as follows. Ovaries were
dissected in ice cold Ringer’s solution (or males were
collected) and frozen at -70°C until use. Ice cold Ripa
buffer (see Harlow and Lane, 1988) was added to frozen
ovaries and the tissue was homogenized in an Eppendorf
tube. Samples were sonicated for 2 min at maximum
power in a cup sonicator and 2 x protein sample buffer
(see Maniatis et al., 1989) was added. Samples were
boiled for 5 min, spun for 10 min in a microfuge and su-
pernatants loaded on a 10% polyacrylamide gel. Western
blots were probed with anti-Grk antibody at a final dilu-
tion of 1:4000. Horseradish peroxidase (HRP)-conjugated
goat anti-rat (Southern Biotechnology) at a final dilution
of I:1000 was used as a secondary antibody. Protein
bands were detected by enhanced chemiluminescence
(ECL) (Amersham). Total protein loading was normal-
ized by probing the blots with a-tubulin (Sigma). For in
vitro protein synthesis we used the TNT T7-Coupled Re-
112 F. Shiru Neumun-Silberberg, T. Schiipbach /Mechanisms of Development 59 (1996) 105-113
ticulocyte Lysate system (Promega) and included the
1.7 kb grk cDNA plasmid (Neuman-Silberberg and
Schiipbach, 1993) as template for the coupled transcrip-
tion/translation reaction.
Lindsley, D.L. and Zimm, G.G. (1992) The Genome of Drosophilu
melunoguster, Academic Press, New York.
Mahowald, A.P. and Kambysellis, M.P. (1980) Oogenesis. In Ash-
burner, M. and Wright, T.R.F. (eds.), The Genetics and Biology
of Drosophilu, Vol. 2D, Academic Press, New York, pp. 141-
225.
Acknowledgements
We would like to thank Elizabeth Gavis, Laura Nilson
and Ann-Marie Queenan for comments on the manuscript
and all members of the Schiipbach and Wieschaus labora-
tories for encouragement and discussions. We are particu-
larly grateful to Joe Goodhouse for instruction and excel-
lent assistance with the confocal microscope. F.S.N.-S.
was supported by a New Jersey Commission on Cancer
Research Postdoctoral Fellowship. This work was sup-
ported by the Howard Hughes Medical Institute and by
the Public Health Service Grant GM40558 to T.S.
Maniatis, T., Fritsch, E.F. and Sambrook, J. (1989) Molecular Cloning;
a Laboratory Manual, 2nd edn., Cold Spring Harbor Laboratory
Press, Cold Spring Harbor, NY.
Manseau, L.J. and Schilpbach, T. (1989) Cuppuccino and spire: two
unique maternal-effect loci required for both the anteroposterior and
dorsoventral patterns of the Drosophila embryo. Genes Dev. 3,
1437-1452.
Montell, D.J. Keshishian, H. and Spradling, A.C. (1991) Laser ablation
studies of the role of the Drosophilu oocyte nucleus in pattern for-
mation. Science 245,29@-293.
References
Bejsovec, A. and Wieschaus, E. (1995) Signaling activities of the Dro-
sophilu wingless gene are separately mutable and appear to be
transduced at the cell surface. Genetics 139, 309-320.
Christerson, L.B. and McKearin, D.M. (1994) orb is required for an-
teroposterior and dorsoventral patterning during Drosophila
oogenesis. Genes Dev. 8.614628.
Christenson, L.B., Lantz, V., Chang, J., Schedl, P. and McKearin, D.M.
(1995) The Orb protein regulates RNA targeting in the Drosophila
ovary. In Lipshitz, H.D. (ed.), Localized RNAs, Springer Verlag.
Heidelberg, Germany, pp. 87-98.
Neuman-Silberberg, F.S. and Schiipbach, T. (1993) The Drosophih
dorsoventral patterning gene gurken produces a dorsally localized
RNA and encodes a TGFa-like protein. Cell 75, 165-174.
Neuman-Silberberg, F.S. and Schiipbach, T. (1994) Dorsoventral axis
formation in Drosophila depends on the correct dosage of the gene
g&en. Development 120,2457-2463.
Peifer, M., Orsulic, S., Sweeton, D. and Wieschaus, E. (1993) A role
for the Drosophib segment polarity gene urmudillo in cell adhesion
and cytoskeletal integrity during oogenesis. Development 118,
1191-1207.
Pokrywka, N.J. and Stephenson, E.C. (1991) Microtubules mediate the
localization of bicoid RNA during Drosophiku oogenesis. Devel-
opment 113,55-66.
Price, J.V., Clifford, R.J. and Schilpbach, T. (1989) The maternal ven-
tralizing locus torpedo is allelic to jtiint little bull, an embryonic
lethal, and encodes the Drosophiku EGF receptor homolog. Cell 56,
1085-1092.
Clark, 1.. Giniger, E., Ruohola-Baker, H., Jan, L.Y. and Jan, Y.N.
(1994) Transient posterior localization of a kinesin fusion protein
reflects anterioposterior polarity of the Drosophila oocyte. Curr.
Biol. 4, 298-300.
Rongo, C. and Lehmann, R. (1996) Regulated synthesis, transport and
assembly of the Drosophilu germ plasm. Trends Genet. 12, 102-
109.
Clifford, R.J. and Schtipbach, T. (1989) Coordinately and differentially
mutable activities of torpedo, the Drosophila melunoguster ho-
molog of the vertebrate EGF receptor gene. Genetics 123, 771-
787.
Roth, S. and Schiipbach, T. (1994) The relationship between ovarian
and embryonic dorsoventral patterning in Drosophila. Development
120,2245-2257.
Gonzalez-Reyes, A., Elliott, H. and St Johnston, D. (1995) Polarization
of both major body axes in Drosophila by gurken-torpedo signal-
ling. Nature 375,654-658.
Roth, S., Neuman-Silberberg, F.S., Barcelo, G. and Schtipbach, T.
(1995) cornichon and the EGF receptor signaling process are neces-
sary for both anterior-posterior and dorsal-ventral pattern formation
in Drosophila. Cell 81, 967-978.
Hake, L.E. and Richter, J.D. (1994) CPEB is a specific factor that me-
diates cytoplasmic polyadenylation during Xenopus oocyte matura-
tion. Cell 79.617-627.
Ruohola, H., Bremer, K.A., Baker, D., Swedlow, J.R., Jan, L.Y. and
Jan, Y.N. (1991) Role of neurogenic genes in establishment of fol-
licle cell fate and oocyte polarity during oogenesis in Drosophih.
Cell 66.433-449.
Harlow, E. and Lane, D. (1988) Antibodies: a Laboratory Manual, Cold
Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Kelley, R.L. (1993) Initial organization of the Drosophitu dorsoventral
axis depends on an RNA-binding protein encoded by the squid
gene. Genes Dev. 7.948-960.
Ruohola-Baker, H., Grell, E., Chou, T.B., Baker, D., Jan, L.Y. and Jan,
Y.N. (1993). Spatially localized Rhomboid is required for estab-
lishment of the dorsal-ventral axis in Drosophila oogenesis. Cell
73,953-965.
Koch, E.A. and Spitzer, R.H. (1983) Multiple effects of colchicine on
oogenesis in Drosophilu: induced sterility and switch of potential
oocyte to nurse-cell development pathway. Cell Tissue Res. 228,
21-32.
Salles. F.J., Lieberfarb, M.E., Wreden C, Gergen, J.P. and Strickland, S.
(1994) Coordinate initiation of Drosophila development by regu-
lated polyadenylation of maternal mRNAs. Science 226, 1996-
1999.
Lane, E.L. and Kalderon, D. (1994) RNA localization along the antero-
posterior axis of the Drosophila oocyte requires PKA-mediated sig-
nal transduction to direct normal microtubule organization. Genes
Dev. 8, 29862995.
Lantz, V., Ambrosio, L. and Schedl, P. (1992) The Drosophila orb
gene is predicted to encode sex-specific germ line RNA-binding
proteins and has localized transcripts in ovaries and early embryos.
Development 115, 75-78.
Schejter, E.D. and Shilo, B.-Z. (1989) The Drosophilu EGF receptor
homolog (DER) gene is allelic to juint little bull, a locus essential
for embryonic development. Cell 56, 1093-I 104.
Schiipbach, T. (1987) Germ line and soma cooperate during oogenesis
to establish the dorsoventral pattern of egg shell and embryo in
Drosophila melunoguster. Cell 49, 699-707.
Schiipbach, T., Clifford, R.J., Manseau, L.J., and Price, J.V.(1991)
Dorso-ventral signal processes in Drosophilu oogenesis. In Gerhart,
.I. (ed.), Cell-Cell Interaction in Early Development, Wiley-Liss,
New York, pp. 163-174.
Lindsley, D. and Grell, R. (1968) Genetic Variations of Drosophila Schiipbach, T. and Wieschaus, E. (1991) Female sterile mutations on
meluno~u~rter, Carnegie Inst. Wash. Publ. 627. the second chromosome of Drosophila me/uno@~ter. II: mutations
F. Shim Neumun-Silberberg, T. Schiipbuch / Mechanisms qf Development 59 (1996) 105-I 13 113
that affect egg chamber and egg morphology. Genetics 129, 1119-
1136.
Serano, T.L., Karhn-McGinness, M. and Cohen, R.S. (1995) The role
offs(l in the localization of the mRNA of the TGFa homolog
gurken within the Drosophila oocyte. Mech. Dev. 51, 183-192.
Spradling, A.C. (1993) Developmental genetics of oogenesis. In Bate,
M. and Martinez-Arias, A. (eds.), The Development of Drosophila
melunoguster, Cold Spring Harbor Laboratory Press, Cold Spring
Harbor, NY, pp. I-70.
St Johnston, D. (1995) The intracellular localization of messenger
RNAs. Cell 81, 161-170.
Wieschaus, E., Marsh, J.L. and Gehring, W. (1978) ,fs(l)KlO, a germ-
line-dependent female sterile mutation causing abnormal chorion
morphology in Drosophdu melunoguster Roux’s Arch. Dcv. Biol.
184,75-82.
Wieschaus, E. (1979) fs(l)KIO, a female sterile mutation altering the
pattern of both the egg coverings and the resultant embryos in Dm-
sophifu. In LeDouarin, N. (ed.). Cell Lineage, Stem Cell and Cell
Differentiation, ElsevierINorth-Holland Biomedical Press, New
York, pp. 291-302.