Available via license: CC BY 4.0
Content may be subject to copyright.
Decreased DNA Repair Efficiency by Loss or Disruption of p53
Function Preferentially Affects Removal of Cyclobutane Pyrimidine
Dimers from Non-transcribed Strand and Slow Repair Sites in
Transcribed Strand*
(Received for publication, November 17, 1999, and in revised form, January 6, 2000)
Qianzheng Zhu‡, Manzoor A. Wani‡, Mohammed El-mahdy‡, and Altaf A. Wani‡§¶储
From the ‡Department of Radiology, §Biochemistry Program, and ¶James Cancer Hospital and Research Institute, Ohio
State University, Columbus, Ohio 43210
The tumor suppressor protein p53 plays a central role
in modulating the cellular responses to DNA damage.
Several recent studies, undertaken with the whole
genomic DNA or full-length gene segments, have shown
that p53 is involved in nucleotide excision repair and it
selectively influences the adduct removal from the non-
transcribed strand in the genome. In this study, we have
analyzed the damage induction at nucleotide resolution
by ligase-mediated polymerase chain reaction and com-
pared the repair of ultraviolet radiation-induced cy-
clobutane pyrimidine dimers within exon 8 of p53 gene
in normal and Li-Fraumeni syndrome fibroblasts as well
as in normal and human papillomavirus 16 E6 and E7
protein-expressing human mammary epithelial cells.
The results demonstrate that (i) loss or disruption of p53
function decreases efficiency of DNA repair, by prefer-
entially affecting the repair of non-transcribed strand
and of intrinsically slow repair sites in transcribed
strand; (ii) mutant p53 protein affects DNA repair, at
least of non-transcribed strand, in a dominant negative
manner; and (iii) pRb does not have an effect on the
repair of DNA damage within transcribed or non-tran-
scribed strand. The overall data suggest that p53 could
regulate excision repair or related events through di-
rect protein-protein interaction.
Mammalian cells have evolved sophisticated DNA repair
mechanisms to overcome DNA damaging hazards that
threaten the integrity of genome (1–4). The most versatile and
thoroughly studied repair system is the excision repair, of
which two major pathways, nucleotide excision repair (NER)
1
and base excision repair, have been identified (5, 6). NER
removes many types of DNA lesions including cyclobutane
pyrimidine dimers (CPDs) by global genomic repair (GGR) and
transcription-coupled repair (TCR) (5–8). It is now well estab-
lished that NER along genome is heterogeneous, CPDs are
more efficiently removed from transcriptionally active genes,
and TCR is generally faster than GGR (2, 9, 10). GGR acts on
the elimination of lesions from non-transcribed strand (NTS)
and transcriptionally inactive genes, whereas TCR removes
lesions from DNA strand transcribed by RNA polymerase II.
In mammalian cells, a variety of cellular responses following
genotoxic exposure may contribute to prevent DNA lesions
from interfering with essential cellular functions. Of those
cellular responses, activation of p53 pathway is well studied
and documented (11, 12). p53 is a critical protein for maintain-
ing genomic stability and homeostasis. It is believed that p53
activation signals the G
1
arrest to delay the transit from G
1
to
S, thus preventing the effects of DNA lesions on vital cellular
functions. Accumulating evidence indicates that p53 plays a
role in DNA repair, especially in GGR (13–17). Viral proteins
that bind p53, causing p53 inactivation or degradation, inter-
fere with p53-regulated DNA repair (14, 18–20). Conceivably,
p53 could be involved in NER by regulating its downstream
genes, which are either related to or actively participate in
NER. For example, Gadd45 and p21
waf1
, which are among the
many p53-regulated proteins, have been shown to interact with
proliferating cell nuclear antigen (21). Additionally, recent ev-
idence has shown that p48, a UV-damaged DNA-binding pro-
tein is transcriptionally regulated by p53, linking p53 to NER
(22). Besides transcriptional activation, p53 may also be di-
rectly involved in repair or repair-related processes. For exam-
ple, in addition to demonstrated p53 binding to insertion/dele-
tion mismatches (23), p53 has also been found to both
physically and functionally interact with p62, XPD, and XPB,
three components of basal transcription factor IIH (10, 24).
Despite a plethora of experimental data, the definitive role of
p53 in NER and the mechanistic basis of its interaction with
repair machinery has not been fully delineated in eukaryotic
systems. Several observations, either supporting a pronounced
effect of p53 for an efficient repair of NTS alone or its involve-
ment also in the repair of transcribed strand (TS), have been
put forth to identify the role of p53 in NER (13–17, 19, 25, 26).
These observations are primarily based upon the assessment of
an average of the repair events within an entire genome, a
specific gene segment, or in some cases an episomally replicat-
ing plasmid within mammalian host cells. Thus, in the present
study we performed the damage analysis at nucleotide resolu-
tion and systematically compared DNA repair of individual
UV-induced CPD sites in normal, p53 mutant (p53-Mut), and
p53 nullizygous (p53-Null) Li-Fraumeni syndrome (LFS) fibro-
blasts, as well as in normal, human papillomavirus (HPV) 16
E6 and E7 protein-expressing human mammary epithelial cells
(HMEC). Our results show that, compared with the efficiency
* This work was supported by National Institutes of Health NIEHS
Grant ES2388. The costs of publication of this article were defrayed in
part by the payment of page charges. This article must therefore be
hereby marked “advertisement” in accordance with 18 U.S.C. Section
1734 solely to indicate this fact.
储To whom correspondence should be addressed: Molecular Carcino-
genesis Laboratory, 103 Wiseman Hall, 400 W. 12th Ave., Columbus,
OH 43210. Tel.: 614-292-9375; Fax: 614-292-7237; E-mail: wani.2@
osu.edu.
1
The abbreviations used are: NER, nucleotide excision repair; CPD,
cyclobutane pyrimidine dimer; LFS, Li-Fraumeni syndrome; HMEC,
human mammary epithelial cell; LMPCR, ligation-mediated polymer-
ase chain reaction; WT, wild type; Mut, mutant; Null, nullizygous;
HPV, human papillomavirus; NTS, non-transcribed strand; TS, tran-
scribed strand; TCR, transcription-coupled repair; GGR, global genomic
repair.
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 275, No. 15, Issue of April 14, pp. 11492–11497, 2000
© 2000 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A.
This paper is available on line at http://www.jbc.org11492
of TS repair, a significantly reduced rate of CPD removal was
observable at all the dipyrimidine target sites of exon 8 NTS in
both p53-Null and p53-Mut LFS cell lines. A close comparison
of the extent of damage removal at 24 and 48 h after UV
irradiation indicated that the repair of NTS was more severely
affected in p53-Mut cells than in p53-Null cells. Relatively
slight decrease in the initial rate of repair within TS, discern-
ible at 24 h after treatment, was fully apparent at the inher-
ently slow repair sites, e.g. codons 286 and 294 of p53 gene.
This selective effect of p53 protein on the slow repair site of TS
was much more clearly demonstrable at the same site in HPV
16 E6-expressing HMEC in which p53 is functionally inacti-
vated by E6 protein. Like LFS cells, HPV 16 E6-expressing
HMEC show a reduced rate of repair of CPDs in the NTS than
in the TS. Both normal HMEC, having wild-type p53 and pRb
proteins, and the isogenic HPV 16 E7-expressing HMEC, hav-
ing wild-type p53 and compromised pRb protein expression,
failed to show the deficiency of CPD repair in NTS as seen
clearly due to the absence of p53 within human cells. Thus,
pRb, another important cell cycle regulatory protein, does not
seem to influence the NER process to a meaningful extent.
MATERIALS AND METHODS
Cell Culture and Treatment—The normal human (p53-WT) fibro-
blasts (OSU-2) were established in culture as described earlier (27).
LFS fibroblast strains, MDAH087 (p53-Mut, harboring a codon 248
single-base substitution) and MDAH041 (p53-Null, harboring a codon
184 frameshift mutation, resulting in premature termination of trans-
lation of p53 protein), both ⬎200 population doubling, post-crisis p53
homozygous cell strains, were kindly provided by Dr. Michael Tainsky
(M. D. Anderson Cancer Center, Austin, TX). These fibroblast cells were
grown in DMEM supplemented with 10% fetal calf serum and antibi-
otics at 37 °C in a humidified atmosphere of 5% CO
2
. Normal HMEC
were established in culture according to Stampfer (28). HPV 16 E6
protein-expressing HMEC 76E6 and HPV 16 E7 protein-expressing
HMEC 76E7 were kindly provided by Dr. Vimla Band (Tufts University
School of Medicine, Boston, MA). These cells were grown in DFCI
medium supplemented with required nutrients and growth factors as
described (29). For experiments to assess DNA damage and repair, the
monolayer cells were grown to confluence in 150-mm dishes and then
placed in serum-free medium for 12 h. Under these growth and main-
tenance conditions, the index of cellular DNA replication, as measured
from changes in the specific radioactivity of DNA pre-labeled with
[
3
H]dThd, does not show significant genome duplication during the test
periods of repair analysis (27). The medium was removed, and cells
were washed with prewarmed phosphate-buffered saline and irradiated
with desired doses of UV (254-nm) light at a rate of 0.5 J䡠m
⫺2
䡠s
⫺1
as
measured by Kettering model 65 radiometer (Cole-Palmer Instrument
Co., Vernon Hill, IL). After irradiation, fresh serum-free medium was
added to cell culture and incubation continued for desired periods.
DNA Isolation and Conversion of CPDs to Ligatable DNA Strand
Breaks—Briefly, after UV exposure, desired incubation periods, and
washes to remove any floating dead cells, the adherent cells were
recovered by trypsinization and immediately lysed for DNA isolation by
a salt precipitation procedure as described (13, 30). The CPDs were
cleaved and converted to single-strand breaks by digestion with T4
endonuclease V (31). The ligation-inhibiting 5⬘-pyrimidine overhang
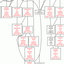
FIG.1.Repair of CPDs in non-transcribed strand of exon 8 of p53 in genomic DNA. Human fibroblasts were irradiated with an UV dose
of 20 J䡠m
⫺2
and DNA isolated at indicated times after irradiation. CPDs in non-transcribed strand of genomic DNA were determined by LMPCR
using upper strand specific primers as described under “Materials and Methods.” Maxam-Gilbert-derived sequencing lanes, lanes C⫹Tand C, are
shown on the left of each autoradiogram. Formation or remaining amount of CPDs is shown at indicated postirradiation time points. Lanes marked
Con represent LMPCR profiles of DNA sample from cells without UV treatment. A, p53-WT normal human fibroblasts; B, p53-Null LFS fibroblasts;
C, p53-Mut LFS fibroblasts.
NER Modulation by p53 11493
was removed by Escherichia coli photolyase (31) (generous gift from Dr.
Aziz Sancar, University of North Carolina, Chapel Hill, NC). DNA was
then recovered and quantitated as described previously (13). The same
amount of DNA (1–2
g) was used for each reaction set of LMPCR.
LMPCR—LMPCR is an extremely sensitive genomic sequencing
method used for the detection of DNA damage (31, 32). DNA, specifi-
cally cleaved at CPDs, was used to create blunt end DNA fragment by
extension of primer specific to p53 with DNA sequenase 2.0. Blunt-
ended DNA was then ligated to a double-strand linker and followed by
amplification with the longer oligonucleotide of the linker and a second
nested p53-specific primer. After 20–21 cycles of polymerase chain
reaction, DNA fragments were size-fractionated on an 8% urea-polyac-
rylamide sequencing gel, electroblotted onto a nylon filter, and hybrid-
ized to a single-stranded p53-specific
32
P-labeled probe generated by
polymerase chain reaction using a third p53-specific primer. The filter
was used to expose a PhosphorImager screen and the individual band
intensities quantitated upon imaging and processing by Imagequant
software (Molecular Dynamics). The filters were also used to expose
Kodak X-Omat film for autoradiography and documentation by image
scanning. LMPCR analysis of each sample was carried out in duplicate,
and results described are from at least three independent experiments.
RESULTS
Loss or disruption of normal p53 function generally results in
decreased efficiency of repair of CPDs of non-transcribed DNA
strand. To compare repair of CPDs in p53-WT containing nor-
mal human fibroblasts, p53-Mut, and p53-Null LFS fibroblasts,
formation and removal of CPDs induced by UV at a dose of 20
J䡠m
⫺2
were mapped by LMPCR, in both strands of exon 8 of p53
from genomic DNA. Under such an exposure condition, no
discernible differences in formation of CPDs in genomic DNA
could be seen in these three cell lines as measured by immuno-
slot blot assay (13). However, repair of CPDs by LMPCR could
be seen at almost all the potential dimer sites of NTS, albeit
with a clearly demonstrable variation in the rates of their
repair. For example, repair at many sites in p53-WT normal
human fibroblasts was apparent after 8 h, followed by an
approximately 30–70% repair after 24 h and 60–95% repair
after 48 h (Fig. 1A, Table I). Higher efficiency of removal of
CPDs was seen at 5⬘side of adjacent pyrimidine sites, e.g.
codons 270, 274, and 289. Consistent with earlier observation
by Tornaletti and Pfeifer (32), repair of CPDs at codon 278,
which is one of the frequently mutated in human skin cancers,
was slower than those of surrounding positions. However, re-
pair of CPDs (5⬘-TCTC^C) at codon 289/290 was also found to
be slower. It may be noted that this dimer is located at the 3⬘
end of four adjacent potential pyrimidine sites.
In p53-Null cells, repair of CPDs at most of the dipyrimidine
sites of NTS was slower compared with that of the same CPDs
at same position in p53-WT cells. Moreover, slow repair sites
FIG.2.Repair of CPDs in transcribed strand of exon 8 of p53
genomic DNA in p53-WT, p53-Null, and p53-Mut LFS fibroblasts.
The LMPCR processing of the sample was as described under “Materi-
als and Methods.” The composite is a representative autoradiogram of
data from several independent experiments.
TABLE I
Repair of CPDs of exon 8 of p53 genomic DNA in normal and LFS fibroblasts
DNA Codon Sequence (5⬘–3⬘)
CPD remaining
a
after
p53-WT p53-Mut p53-Null
24 h 48 h 24 h 48 h 24 h 48 h
Non-transcribed %
270 T∧TT 22 6 65 60 58 38
TT∧T 4912 73606044
274/275 GTT∧T 6848 80566042
276/277 C∧TG 22 11 100 53 50 22
277/278(M) T∧CCT 36 34 100 53 65 40
278(M) C∧CT 32 18 100 97 65 40
289 C∧TC 10 9 100 80 34 20
CT∧C 17 10 100 80 54 22
289/290 CTC∧C 41 15 100 84 60 30
Transcribed 285/286(M) C∧CTC 44 23 62 31 40 40
286(M) TT∧C 4130 55315232
291 CT∧T 3018 40204722
292 TT∧T 10 8 34 8 30 14
293 C∧CC 32 17 47 20 34 8
294(M) CT∧C 3620 75504220
a
Repair rates were measured at mutation hotspots (M) and at various surrounding sites. Repair at each position is described as average
percentage derived from time versus repair plots of three independent experiments. Percentage of CPDs remaining was calculated from band
intensities at 24 and 48 h in reference to the band intensity at 0 h and normalized for any intensity observed at same sites in control lanes.
NER Modulation by p5311494
were more prominently affected (Fig. 1, Aand B, Table I). For
example, dimer C^CT, at codon 278, was 68% repaired at 24 h
and 82% repaired at 48 h in p53-WT cells, whereas in p53-Null
cells, there was only 35% and 60% repair observed at these
sites within 24 and 48 h, respectively. Repair of CPDs (5⬘-
TCTC^C) at codon 289/290 was also drastically affected by the
loss of p53 function (Fig. 1B). These observations were further
confirmed by comparison of normal HMEC with HPV 16 E6
protein-expressing HMEC. In the case of 76E6 HMEC, in
which p53 protein is degraded by E6 protein-mediated ubiq-
uitin proteolysis pathway (33), the overall p53 modulation of
DNA repair events appeared exactly like that of p53-Null fi-
broblast cells described above (redundant data not shown).
Among the cell lines tested, repair of CPDs in NTS was most
dramatically affected in p53-Mut fibroblasts (Fig. 1Cand Table
I). In this cell line, repair of CPDs at all dipyrimidine sites was
significantly slow. Approximately 80–100% of CPDs remained
after 24 h, and 50–100% of CPDs remained 48 h after UV
irradiation. These results, in conjunction with the data from
p53-Null cells, indicate that mutant p53 protein affects DNA
repair in a dominant negative manner. This would seem to
suggest that p53 protein regulates DNA repair, at least of the
non-transcribed strand, by direct protein-protein interaction.
Such a dominant effect could be exerted by interaction with the
proteins of NER assembly or by altered transcription of pro-
teins essential for optimal assembling and targeting of the
damage recognition complex.
At the position of one potential dipyrimidine site, (5⬘-
C^CTCACC) at codon 295, an abnormal signal was distinctly and
reproducibly detected in p53-Null and p53-Mut cells, but not in
p53-WT cells (Fig. 1, A–C). Accordingly, a distinct LMPCR gen-
erated band could be seen in the control unirradiated sample
lane. We surmise that this band could not arise due to nonspecific
or enzymatic cleavage as the signal gradually decreased between
FIG.3.Repair of CPDs in transcribed
strand of exon 8 of p53 genomic DNA in
normal, HPV 16 E6 and E7 protein-ex-
pressing HMEC. A, normal HMEC; B,
HPV 16 E6 protein-expressing HMEC,
76E6; C, HPV 16 E7 protein-expressing
HMEC, 76E7. The LMPCR processing of the
sample was as described under “Materials
and Methods.”
NER Modulation by p53 11495
4- and 48-h time intervals following irradiation. According to the
nature of LMPCR, an assay specialized for detecting nicks in
individual DNA strand, this signal could only represent a specific
DNA strand break at this site; due to its occurrence within DNA
topoisomerase consensus sequence, it could be the result of an
arrested cleavable complex. Surprisingly, such an abnormal
DNA break signal was also found in HPV 16 E6-expressing
HMEC, but not in normal or in HPV 16 E7-expressing HMEC.
This further confirmed that the DNA strand break at this codon
site is specific and only appears in cells that are rendered defi-
cient for normal p53 function. Interestingly, exposure of cells to
UV seemed to induce the repair of this DNA strand break, as was
evident from time-dependent gradual disappearance of the band
at this site.
Loss or Disruption of Normal p53 Function Affects the Repair
of CPDs at Slow Repair Sites of Transcribed DNA Strand—To
examine the effects of loss of normal p53 function on the repair
of CPDs within TS, repair of CPDs in TS of exon 8 of p53 gene
was mapped by LMPCR (Fig. 2). As expected, removal of CPDs
from TS was generally faster than from NTS, albeit with a
clearly discernible site-specific variation in the removal of
CPDs at individual dipyrimidine sites. Consistent with earlier
observation (32), repair of CPDs in p53-WT cells at codons 286
and 294 was seen to be slower than that of surrounding posi-
tions. A p53-dependent decreased repair of CPDs was found at
several sites of exon 8 in both p53-Null and p53-Mut LFS cells
at 24 h after UV irradiation. The intrinsically slow repair
dipyrimidine sites, e.g. at codons 286 and 294, were preferen-
tially affected by the absence of normal wild type function than
surrounding CPD sites (Fig. 2 and Table I). The intrinsically
slow repair is suggested to be the basis for mutational predis-
position of these p53 gene sites in human cancers (32). An
absence of p53 function would be expected to further exacer-
bate the cellular instability through decreased repair of exog-
enously or endogenously induced DNA damage.
Since the effect of loss of p53 function on TCR has not been
fully resolved, we extensively mapped repair of CPDs in TS of
exon 8 of p53 gene in normal, isogenic HPV 16 E6 and E7
protein-expressing HMECs. It may be noted that HPV 16 E7
protein selectively activates ubiquitin proteolysis pathway
causing degradation of pRb protein, while it stabilizes p53
protein (29). The data shows that the repair of CPD was faster
in HMEC than fibroblast, and dimers at most of the sites were
quantitatively removed within 48 h after UV treatment. Fur-
thermore, unlike normal fibroblasts, slow repair of CPDs
within sites like codon 286 and 294 was not very obvious in
either the normal or HPV 16 E7 protein-expressing HMEC
(Fig. 3, Aand C). Thus, p53-expressing cells appear to have
normal NER despite the absence of a functional pRb protein.
On the other hand, a clearly visible slower repair at the same
sites was found in p53-compromised HPV 16 E6 protein-ex-
pressing HMEC (Fig. 3Band Table II). This observation fur-
ther confirmed that loss of functional p53 affects the removal of
CPDs from TS by TCR and preferentially affects removal of
CPDs at slow repair sites.
DISCUSSION
The biochemical mechanisms of NER involve damage recog-
nition and open complex formation by factors such as XPA,
RPA, XPC, and transcription factor IIH, dual incision of the
damaged DNA strand by endonucleases XPF-ERCC1 and XPG,
repairsynthesismediated by a proliferating cell nuclearantigen-
dependent DNA polymerase and ligation of newly synthesized
DNA strand. The precise reaction mechanisms of NER have
recently been established to a significant extent (for review, see
Refs. 1–6). Nonetheless, how NER is regulated or connected to
other cellular functions still remains to be explored. Several
investigators have examined the involvement of p53 in regula-
tion of NER. Different systems and approaches have been used
for assessment of DNA repair. Immunoassay was mostly used
for direct assessment of GGR, whereas endonuclease-sensitive
site assay was used for examination of strand-specific repair. It
is becoming clear that functional p53 is required for efficient
GGR (13–17). However, due to different views being supported
by various TCR studies (14–17, 19, 26), precise nature of p53
participation in TCR still remains unclear. In this study, we
provide the first detailed analysis of effects of loss of functional
p53 on the removal of CPDs in both DNA strands at nucleotide
resolution. First, the results confirmed that functional p53 is
required for efficient GGR (13, 15, 16, 19). Furthermore, the
results show that expression of mutant p53 protein more sig-
nificantly affects removal of CPDs from non-transcribed strand
than loss of p53 protein, indicating that mutant p53 protein
affects GGR in a dominant negative manner. The results also
show that p53 is involved in TCR and repair of CPDs at slow
repair sites is the first target to sustain meaningful effects due
to the loss of p53 function.
In a recent review, Mckay et al. (17) have strongly argued
that p53 plays a definitive role in TCR. The difference of ob-
servations by various laboratories may be the result of different
assays used to detect strand-specific repair. Using the normal
endonuclease-sensitive site assay, we too were unable to dem-
onstrate any detectable differences in TCR between normal
and LFS fibroblasts as well as between normal and HPV 16 E6
protein-expressing HMEC for both UV- and benzo[a]pyrene
diol epoxide-induced DNA damage.
2
Repair differences be-
tween cells did not become pronounced and meaningful until
full mapping of repair of CPDs in normal, HPV 16 E6 and E7
protein-expressing HMEC was conducted within the same gene
2
Q. Zhu, M. A. Wani, M. El-mahdy, and A. A. Wani, unpublished
results.
TABLE II
Repair of CPDs of exon 8 of p53 genomic DNA in normal, HPV 16 E6 and E7 protein-expressing HMEC
DNA Codon Sequence (5⬘–3⬘)
CPD remaining after
HMEC 76E6
a
76E7
a
24 h 48 h 24 h 48 h 24 h 48 h
Transcribed %
285/286(M) TTC^C 5 1 30 18 10 2
286(M) TT^C 14 2 40 20 8 0
T^TC 26 6 50 16 20 0
287 CT^C 16 0 24 6 12 2
291 CT^T 10 1 13 0 7 0
292 TT^T 16 2 35 10 16 3
293 C^CC 8 0 40 30 14 2
294(M) CT^C 10 0 40 22 12 1
a
76E6 and 76E7 represent HPV 16 E6 and E7 protein-expressing HMEC, respectively. Extent of repair at each site was quantitated at indicated
times as described in Table I.
NER Modulation by p5311496
segment by LMPCR. The reasons are that (i) LMPCR is a much
more powerful assay in demonstrating variations of repair
along particular sites and stretches of specific gene sequences,
(ii) fully discernible differences were visualized mainly during
the initial stages of repair, i.e. before 24-h time points, and (iii)
slow repair sites were more prominently subjected to the influ-
ence exerted by the loss of p53 function.
Several investigations suggest that p53-regulated gene prod-
ucts may participate or be associated with NER. Using host
reactivation assay, it has been shown that UV- and heat shock-
inducible NER is p53-dependent (14, 17). More recently, it has
been shown that the expression of p53-downstream genes, e.g.
p48 gene, was dependent on p53 and involved in GGR (22). It
should be noted that p48 has been suggested to have a role in
the repair of DNA in chromatin and damage recognition. If
effects of loss of function in p53-Null LFS fibroblasts and HPV
16 E6 protein-expressing HMEC on NER reflect requirements
of p53-activated downstream genes in NER, dominant negative
effects of mutant p53 protein on DNA repair of non-transcribed
strand may reflect direct participation of p53 in NER. It has
been shown that p53 binds to three components of basal tran-
scription factor: p62, XPD, and XPB (10). Furthermore, XPB
and XPD cells are deficient in repair of non-transcribed DNA
and inefficiently repair the transcribed strand including se-
quences near the transcription start site (34). However, contact
with these factors may not be the only mechanism by which p53
directly participates in NER. Given that mutant p53 protein
negatively regulates NER by binding XPB and XPD, TCR
should be the first target. However, dominant negative effects
of mutant p53 protein on DNA repair of non-transcribed strand
were more distinctly observed. In fact, wild-type rather than
mutant p53 protein has been shown to inhibit DNA helicase
activity of XPD and XPB (35). In search for the components of
NER complex that could be interacting in vivo with p53, we
have found that recognition of UV-induced damage links p53
pathway to NER and that HHR23A is involved in regulating
transcriptional activity of p53.
3
Interestingly, it has been
shown that XPC protein, complexed with HHR23A and
HHR23B, also plays a role in damage recognition and chroma-
tin unfolding (37). It seems that p53 regulates the early steps of
damage recognition of NER or chromatin unfolding during
NER processes through both transcription activation and pro-
tein-protein interaction. However, no experimental evidence
shows that p53 protein recognizes or binds UV-induced CPDs.
In eukaryotic cells, genomic DNA is wrapped around histone
octamers forming nucleosomes, which are the repeating units
of chromatin. Proteins involved in cellular processes, such as
DNA replication, transcription, and DNA repair, require access
to DNA within chromatin structural hierarchy. Heterogeneity
of NER may partially reflect the accessibility of damaged DNA
to NER components. In support of this, very fast repair has
been seen in both DNA strands near the transcription initia-
tion site (38). It has also been shown that several sequence
positions in 5⬘-flanking region of the tRNA
val
gene, which lacks
TCR, were also repaired more efficiently than the gene itself
(38, 39). In the case of a run of potential CPD sites in the
non-transcribed DNA strand, if each DNA repair event in GGR
is considered as an independent event, as suggested by its
biochemical mechanisms, there is no reason for higher effi-
ciency of CPD removal at 5⬘end. This is only possible if TCR or
transcription somehow helped damaged DNA to become more
accessible at 5⬘end. Thus, besides transcription coupling, ac-
cessibility of damaged DNA contributes a very important pa-
rameter to the heterogeneity of NER. In this regard, p53 may
regulate NER through modulating accessibility of damaged
DNA rather than damage recognition. One would expect that
CBP/p300, a p53 coactivator that has been shown to have
acetyltransferase activity (36), could also be involved in p53-
regulated DNA repair. Investigation of such DNA repair par-
ticipating principles should become an area of active interest in
the near future.
Acknowledgments—We are grateful to Dr. Aziz Sancar for providing
photolyase enzyme and John Croyle for assistance with high resolution
image scanning.
REFERENCES
1. Hanawalt, P. C. (1998) Mutat. Res. Fundam. Mol. Mech. Mutagen. 400,
117–125
2. Petit, C., and Sancar, A. (1999) Biochimie 81, 15–25
3. Hoeijmakers, J. H. J., and Bootsma, D. (1994) Nature 371, 654–655
4. Lindahl, T., Karran, P., and Wood, R. D. (1997) Curr. Opin. Genet. Dev. 7,
158–169
5. Wood, R. D. (1997) J. Biol. Chem. 272, 23465–23468
6. Mullenders, L. H. (1998) Mutat. Res. DNA Repair 409, 59–64
7. Drapkin, R., Sancar, A., and Reinberg, D. (1994) Cell 77, 9–12
8. Hanawalt, P. C., Donahue, B. A., and Sweder, K. S. (1994) Curr. Biol. 4,
518–521
9. Tornaletti, S., and Hanawalt, P. C. (1999) Biochimie 81, 139–146
10. Frit, P., Bergmann, E., and Egly, J.-M. (1999) Biochimie 81, 27–38
11. Oren, M., and Rotter, V. (1999) Cell. Mol. Life Sci. 55, 9–11
12. Agarwal, M. L., Taylor, W. R., Chernov, M. V., Chernova, O. B., and Stark,
G. R. (1998) J. Biol. Chem. 273, 1–4
13. Wani, M. A., Zhu, Q. Z., El-mahdy, M., and Wani, A. A. (1999) Carcinogenesis
20, 765–772
14. Smith, M. L., Chen, I.-T., Zhan, Q., O’Connor, P. M., and Fornace, A. J., Jr.
(1995) Oncogene 10, 1053–1059
15. Ford, J. M., and Hanawalt, P. C. (1995) Proc. Natl. Acad. Sci. U. S. A. 92,
8876–8880
16. Ford, J. M., and Hanawalt, P. C. (1997) J. Biol. Chem. 272, 28073–28080
17. McKay, B. C., Francis, M. A., and Rainbow, A. J. (1997) Carcinogenesis 18,
245–249
18. Ford, J. M., Baron, E. L., and Hanawalt, P. C. (1998) Cancer Res. 58, 599–603
19. Prost, S., Ford, J. M., Taylor, C., Doig, J., and Harrison, D. J. (1998) J. Biol.
Chem. 273, 33327–33332
20. Jia, L., Wang, X. W., and Harris, C. C. (1999) Int. J. Cancer 80, 875–879
21. Smith, M. L., Chen, I. T., Zhan, Q., Bae, I., Chen, C.-Y., Gilmer, T. M., Kastan,
M. B., O’Connor, P. M., and Fornace, A. J., Jr. (1994) Science 266,
1376–1380
22. Hwang, B. J., Ford, J. M., Hanawalt, P. C., and Chu, G. (1999) Proc. Natl.
Acad. Sci. U. S. A. 96, 424– 428
23. Lee, S., Elenbass, B., Levine, A. J., and Griffith, J. (1995) Cell 81, 1013–1020
24. Wang, X. W., Forrester, K., Yeh, H., Feitelson, M. A., Gu, J.-R., and Harris,
C. C. (1994) Proc. Natl. Acad. Sci. U. S. A. 91, 2230–2234
25. McKay, B. C., Ljungman, M., and Rainbow, A. J. (1999) Carcinogenesis 20,
1389–1396
26. Mirzayans, R., Enns, L., Dietrich, K., Barley, R. D. C., and Paterson, M. C.
(1996) Carcinogenesis 17, 691–698
27. Venkatachalam, S., Denissenko, M. F., and Wani, A. A. (1995) Carcinogenesis
16, 2029–2036
28. Stampfer, M. R. (1985) J. Tissue Culture Methods 9, 107–115
29. Wazer, D. E., Liu, X.-L., Chu, Q., Gao, Q., and Band, V. (1995) Proc. Natl. Acad.
Sci. U. S. A. 92, 3687–3691
30. Miller, S. A., Dykes, D. D., and Polesky, H. F. (1988) Nucleic Acids Res. 16,
12–15
31. Tornaletti, S., and Pfeifer, G. (1996) in Technologies for Detection of DNA
Damage and Mutations (Pfeifer, G. P., ed) pp. 199–210, Plenum Press, New
York
32. Tornaletti, S., and Pfeifer, G. P. (1994) Science 263, 1436–1438
33. Scheffner, M., Werness, J. M., Huibregtse, J. M., Levine, A. J., and Howley,
P. M. (1990) Cell 63, 1129–1136
34. Tu, Y., Bates, S., and Pfeifer, G. P. (1997) J. Biol. Chem. 272, 20747–20755
35. Wang, X. W., Yeh, H., Schaeffer, L., Roy, R., Moncollin, V., Egly, J.-M., Wang,
Z., Friedberg, E. C., Evans, M. K., Taffe, B. G., Bohr, V. A., Weeda, G.,
Hoeijmakers, J. H. J., Forrester, K., and Harris, C. C. (1995) Nat. Genet. 10,
188–195
36. Chakravarti, D., Ogryzko, V., Kao, H.-Y., Nash, A., Chen, H., Nakatani, Y.,
and Evans, R. M. (1999) Cell 96, 393–403
37. Baxter, B. K., and Smerdon, M. J. (1998) J. Biol. Chem. 273, 17517–17524
38. Tu, Y., Tornaletti, S., and Pfeifer, G. P. (1996) EMBO J. 15, 675–683
39. Dammann, R., and Pfeifer, G. P. (1997) Mol. Cell. Biol. 17, 219–229
3
Q. Z. Zhu, M. El-mahdy, M. A. Wani, and A. A. Wani, submitted for
publication.
NER Modulation by p53 11497